BSSeq Tools
performance evaluation of bioinformatic tools for bisulfite sequencing in animal methylomes
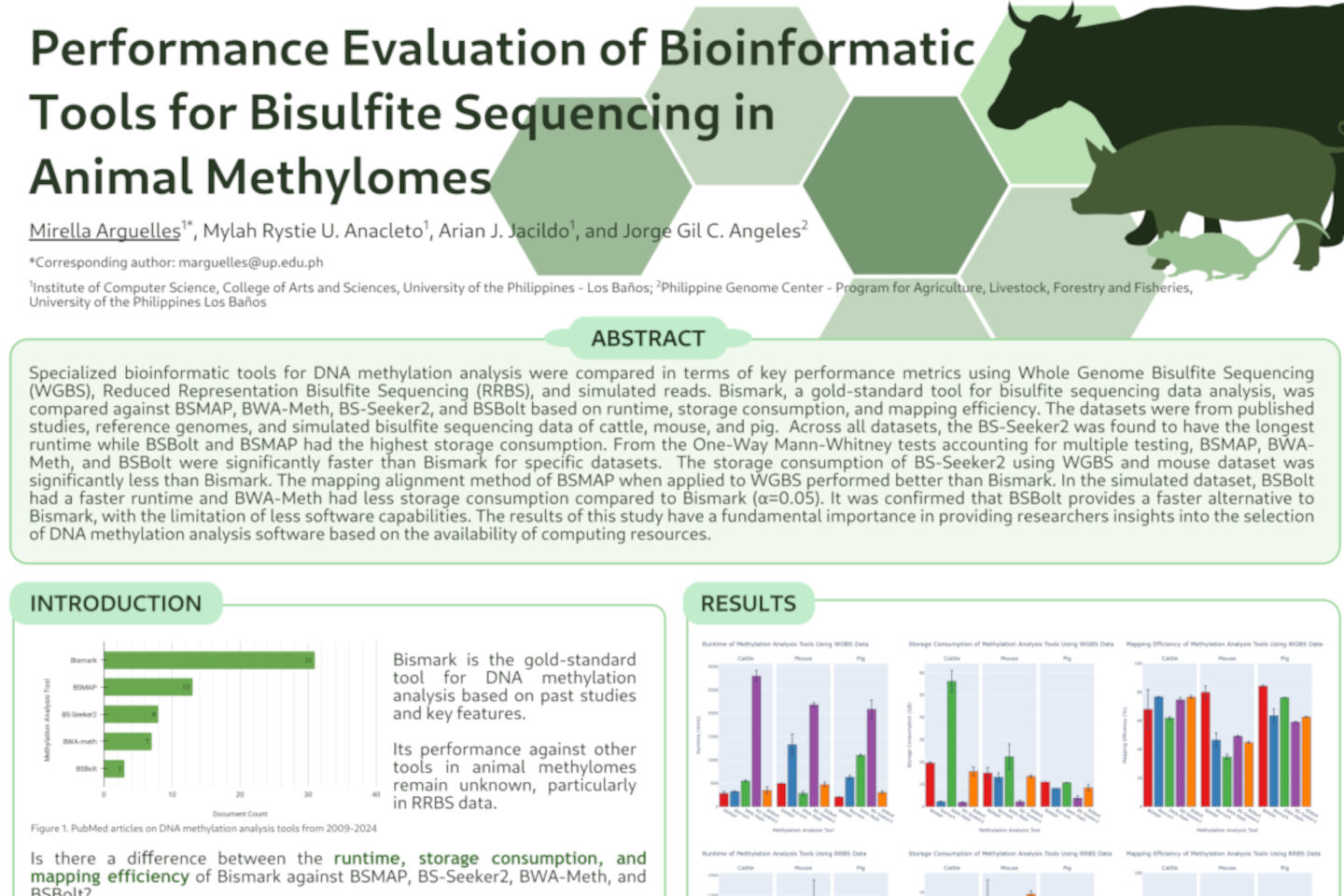
Here, I compared performance metrics of bioinformatic tools in bisulfite sequencing. Whole genome bisulfite sequencing, reduced representation bisulfite sequencing, and simulated reads of cattle, mouse, and pig were used. [Link to project]
Technologies used: Shell scripting, Python, Sherman, Bismark, BSMAP, BSSeeker2, BWA-Meth, BSBolt, Trim Galore, FastQC, Fastp